At Nutricia, we have 120 years of experience in early life nutrition, with over 50 years spent investigating GI function. Our research has enabled us to develop a deep understanding of the impact that early life nutrition can have on the developing GI system, particularly on the composition of the gut microbiota.
Find out more about our working within the broader field of gastrointestinal health and immune health.
Our research programme focuses on understanding how early life events (e.g. mode of delivery, antibiotics) and nutrition (e.g. human milk) influence gut microbial colonisation and development, and gut (barrier) maturation in infants. We also study how different diseases are associated with microbial imbalance and how the microbiota can be modulated through nutrition. In addition, we are also committed to increasing our knowledge on around metabolic load, bioavailability, and kinetics of micro- and macro-nutrients.
Our research enables us to develop optimal nutrition that supports a healthy development of the digestive and metabolic systems.
Some examples of our unique technologies
At Danone Research & Innovation, we are always looking for ways to improve and innovate and, where possible, incorporate state-of-the-art technologies within our research programmes. Two great examples of these are the TIM-1 (TNO Gastro-Intestinal Model) and SHIME (Simulator of Human Intestinal Microbial Ecosystem).
TIM-1 (TNO Gastro-Intestinal Model)
Many of the key benefits of our nutritional products are delivered via mechanisms that occur in the GI system. It is therefore of central importance that we understand how these mechanisms work, which is where the TIM-1 system has proved so valuable. Developed in a collaborative project by TNO and Danone Research & Innovation starting in the 1993, TIM-1 is able to mimic the conditions found in the upper GI tract of infants.Therefore it enables us to study and gain insights in the fate of human milk or formula components after ingestion by the infant in an easy, accessible and reproducible way.
TIM-1 is a computer-controlled model of the stomach and small intestine
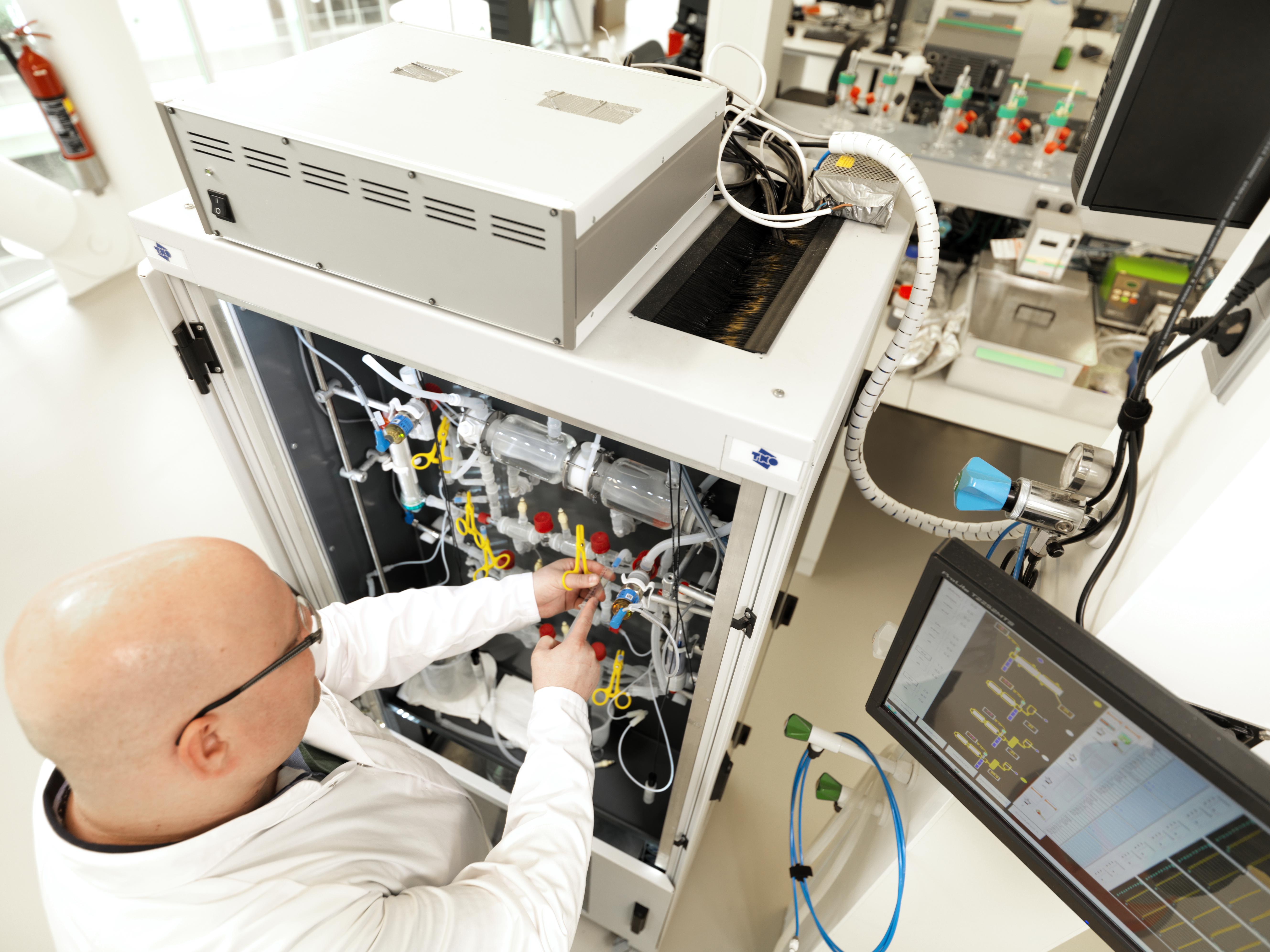

TIM-1 is linked up to a computer system that allows us to control the conditions within the ‘GI tract’ via a series of pumps and sensors allowing us to, for example, secrete digestive enzymes or control pH flow of the food. The parts of the system mirror the human GI tract. There is close resemblance of the dynamic biochemical and physical conditions in the infant GI tract. Therefore no ethical constraints and low variability compared to in vivo trials. It allows sampling at multiple sites of the GI tract.
Once food is entered into the ‘GI tract’, the experiment starts and is carried out in real time (approximately six hours) i.e. different digestive enzymes are released from different part of the system accordingly. We are able to remove samples from a number of points in order to analyse the degree of breakdown or the amount of compound that is still stable. So we can assess the digestion of macronutrients, the bioaccessibility of micronutrients and the stability of bioactive components of human milk and infant formula in the upper GI tract
By the push of a button, we can let a digestion process occur that is close to the in vivo situation as the conditions found in an infant.
One example of its use relates investigating the survival of lactate enzymes, the active compound in our fermented infant formula. The formula was introduced into the ‘stomach’ of the system before letting the experiment run. After six hours, we were able to collect samples from the ileal compartment (the final section of the small intestine) in order to find out how much of the lactase activity was still present.
SHIME (Simulator of Human Intestinal Microbial Ecosystem)
SHIME is an in vitro model that provides a dynamic simulation of the human gut microbial community in the different section of the colon.
Effects and interaction between our nutritional products and the gut microbiota can be investigated using the simulation.
The simulation of the gut microbiota is based on the human fecal sample, collected from a specific age group of interest, inoculated into the colon of the system. The microbial community is then allowed to adapt to the in vitro environment, closely mimicking the physiological conditions of human.
The dynamic simulation of the gut microbiota provides a good platform for investigating the different intervention using our products or screening for new potential candidates of prebiotics and probiotics.

Proof-of-concept studies, such as pathogen infection and antibiotic effects on gut health, can be challenging and difficult to perform in animal models or human clinical trials. This can be made possible using the SHIME.
Our Baby SHIME is the first of its kind in the world and is used to add valuable insight into studies of the gut microbiota of infant and young children.
Thanks to the TIM and SHIME models which enable us to further our knowledge of the fascinating gastro-intestinal system.


